Layered PyMOL renders#
Portein composes PyMOL ray-traced images by stacking layers — each layer is its own PyMOL representation rendered independently, then alpha-composited together with user-controlled transparencies.
Any setting valid for pymol.cmd.set(...) can be passed via
portein.PymolConfig’s pymol_settings dict.
import tempfile
from pathlib import Path
import yaml
from biotite import structure as struct
from biotite.structure import io as bio
from PIL import Image
import portein
The package ships some default PyMOL settings under configs/:
with open("../../configs/pymol_settings.yaml") as f:
pymol_settings = yaml.safe_load(f)
pymol_settings
{'ambient': 0.5,
'antialias': 2,
'cartoon_discrete_colors': True,
'cartoon_fancy_helices': True,
'cartoon_sampling': 20,
'depth_cue': False,
'hash_max': 300,
'light_count': 1,
'ray_opaque_background': False,
'ray_shadows': False,
'ray_texture': 0,
'ray_trace_disco_factor': 1,
'ray_trace_fog': False,
'ray_trace_gain': 0,
'ray_trace_mode': 1,
'specular': False,
'surface_quality': 2}
output_dir = Path(tempfile.mkdtemp())
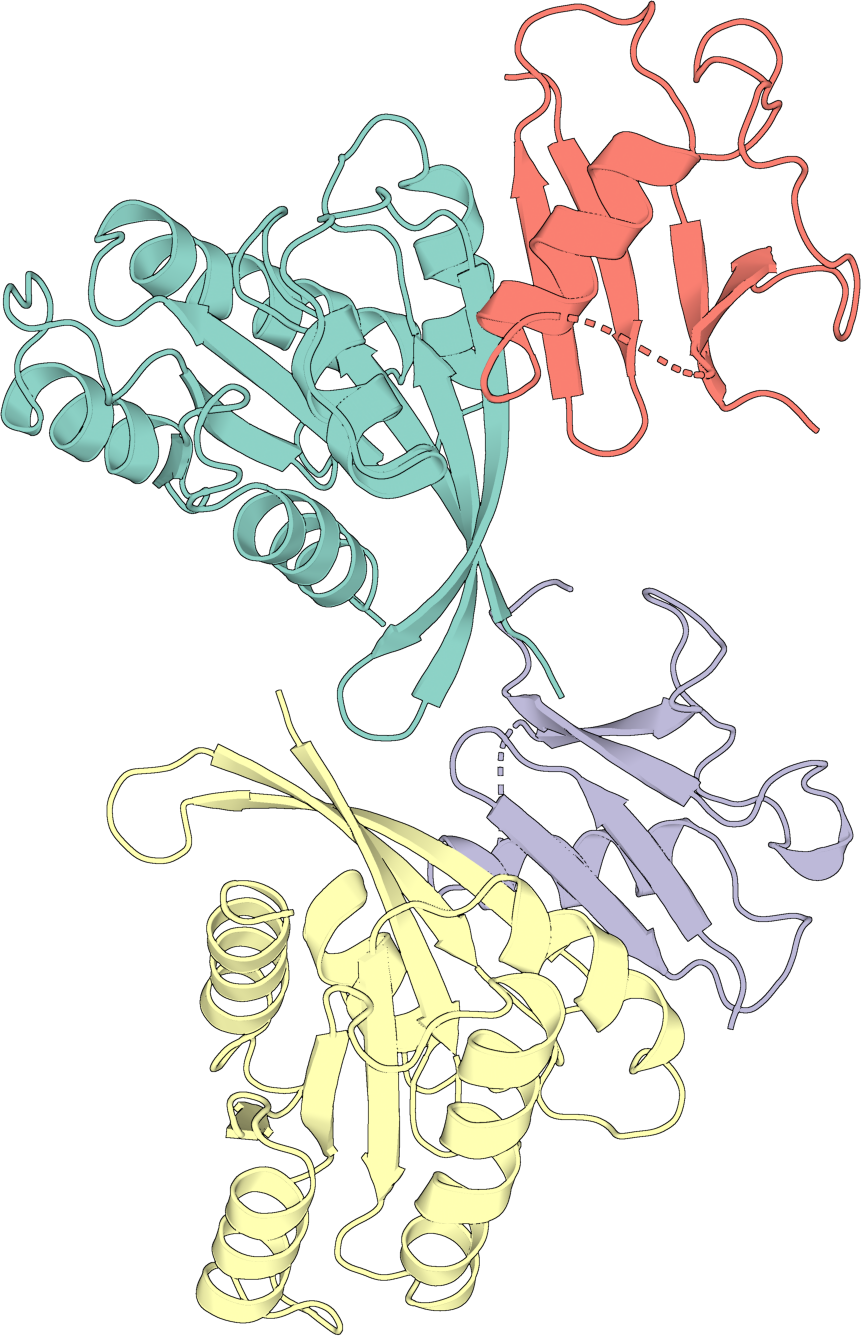
Simple cartoon#
A single-layer cartoon, colored by chain. rotate=True orients the structure
before rendering.
protein_config = portein.ProteinConfig(
pdb_file="../_data/7lc2.pdb",
rotate=True,
width=1000,
chain_colormap="Set3",
output_prefix=str(output_dir / "7lc2_simple"),
)
pymol = portein.Pymol(
protein=protein_config,
layers=[portein.PymolConfig(representation="cartoon", pymol_settings=pymol_settings)],
)
image_file = pymol.run()
portein.crop_to_content(Image.open(image_file))
/home/runner/work/portein/portein/portein/rotate.py:23: NumbaPerformanceWarning: np.dot() is faster on contiguous arrays, called on (Array(float64, 1, 'A', False, aligned=True), Array(float64, 1, 'A', False, aligned=True))
m = find_best_projection(coords)
/home/runner/work/portein/portein/portein/rotate.py:23: NumbaPerformanceWarning: np.dot() is faster on contiguous arrays, called on (Array(float64, 2, 'C', False, aligned=True), Array(float64, 1, 'A', False, aligned=True))
m = find_best_projection(coords)
/home/runner/work/portein/portein/portein/rotate.py:28: NumbaPerformanceWarning: np.dot() is faster on contiguous arrays, called on (Array(float64, 2, 'A', False, aligned=True), Array(float64, 2, 'F', False, aligned=True))
matrix = rotate_to_maximize_bb_height(coords[:, :2]) @ matrix
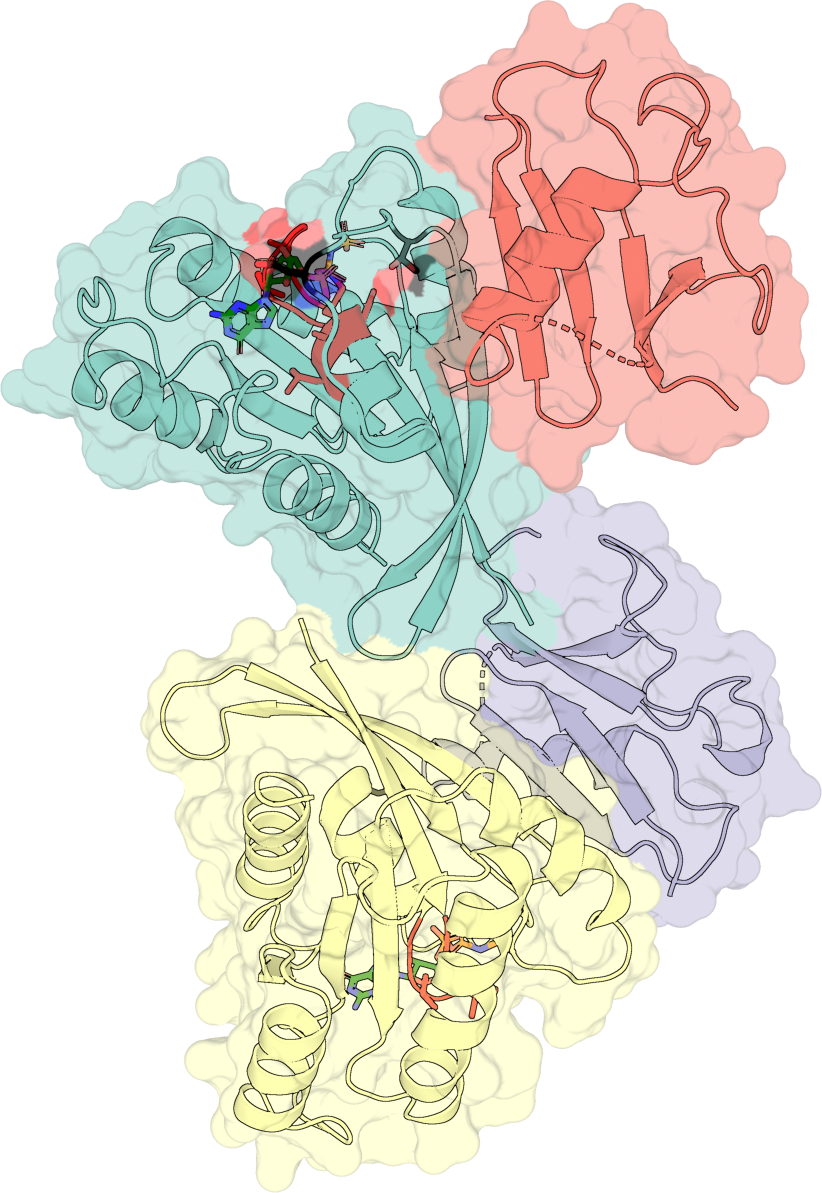
Four-layer composite#
Layer the surface (50% opacity), the cartoon, sticks for highlighted
residues, and sticks colored green for the bound ligand. The
selection="highlight" keyword picks up the residues passed via
highlight_residues on the ProteinConfig.
protein_config = portein.ProteinConfig(
pdb_file="../_data/7lc2.pdb",
rotate=True,
chain_colormap="Set3",
highlight_residues={
"A": {"black": [30, 35], "red": list(range(10, 20))},
"B": {"black": [25], "red": list(range(10, 16))},
},
width=1000,
output_prefix=str(output_dir / "7lc2"),
)
layers = [
[
portein.PymolConfig(representation="surface", pymol_settings=pymol_settings, transparency=0.5),
portein.PymolConfig(representation="cartoon", pymol_settings=pymol_settings),
portein.PymolConfig(representation="sticks", pymol_settings=pymol_settings, selection="highlight"),
portein.PymolConfig(
representation="sticks",
pymol_settings=pymol_settings,
selection="resn GNP",
color="green",
),
],
]
pymol = portein.Pymol(protein=protein_config, layers=layers, buffer=10)
image_file = pymol.run()
portein.crop_to_content(Image.open(image_file))
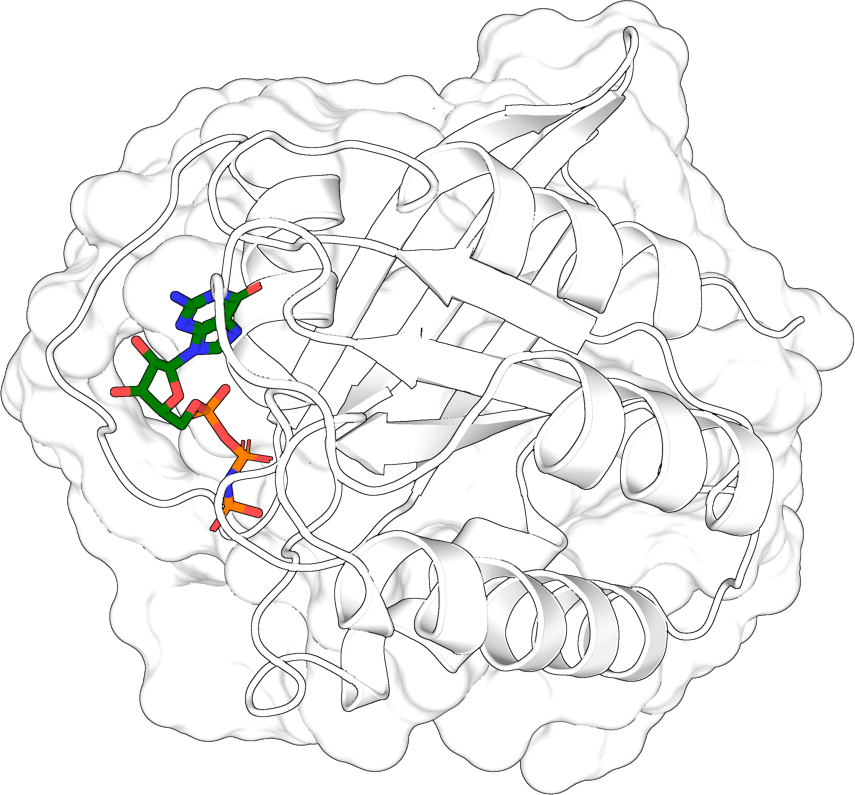
Zoom on a ligand pocket#
Pick out the residues near a bound ligand, rotate to maximize the pocket’s projection, save only the proximal chains, and render with a translucent surface.
import numpy as np
pdb = portein.read_structure("../_data/7lc2.pdb")
ligand = pdb[(pdb.chain_id == "A") & (pdb.res_name == "GNP")]
mask = struct.CellList(pdb, 6).get_atoms(ligand.coord, 6, as_mask=True).any(axis=0)
ligand_pocket = pdb[mask]
proximal_chains = struct.get_chains(ligand_pocket)
# Orient to best-show the pocket
rotation, translation = portein.get_best_transformation(ligand_pocket.coord.astype(np.float64))
pdb_oriented = struct.rotate(struct.translate(pdb, translation), rotation)
# Save only the proximal chains
pocket_path = output_dir / "7lc2_ligand.pdb"
bio.save_structure(str(pocket_path), pdb_oriented[np.isin(pdb_oriented.chain_id, proximal_chains)])
protein_config = portein.ProteinConfig(
pdb_file=str(pocket_path),
rotate=False,
chain_colormap="white",
width=1000,
output_prefix=str(output_dir / "7lc2_pocket"),
)
layers = [
# Surface as its own layer
portein.PymolConfig(representation="surface", pymol_settings=pymol_settings, transparency=0.3),
# Cartoon + ligand sticks share one ray-trace so they depth-interleave.
[
portein.PymolConfig(representation="cartoon", pymol_settings=pymol_settings),
portein.PymolConfig(
representation="sticks",
pymol_settings=pymol_settings,
selection="(chain A and resn GNP)",
color="green",
),
],
]
pymol = portein.Pymol(protein=protein_config, layers=layers)
image_file = pymol.run()
portein.crop_to_content(Image.open(image_file))
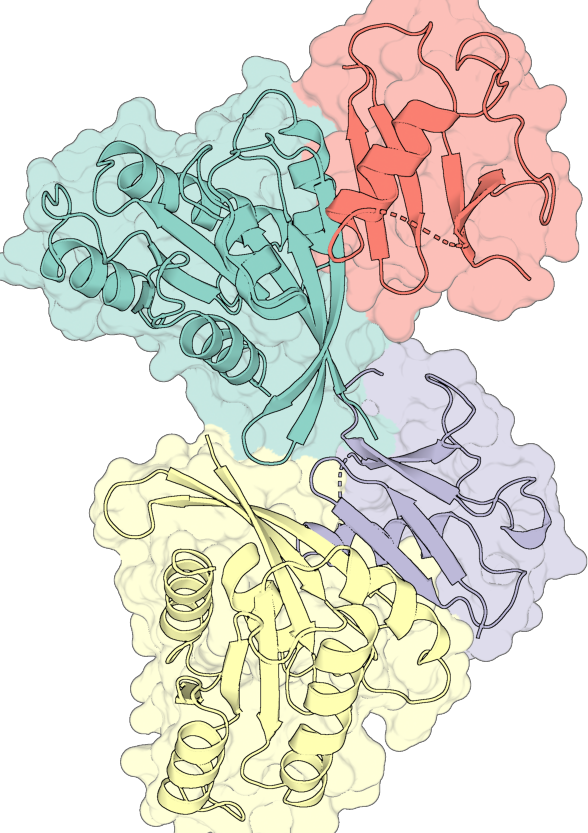
Layer compositing#
Each entry in layers is either a bare PymolConfig or a nested list of PymolConfigs, and that distinction controls how the entry’s geometry composites with the rest:
A bare
PymolConfigis rendered as its owncmd.ray()pass, and itstransparencybecomes a flat 2D PIL alpha across the layer’s PNG before alpha-compositing. This is the right tool for elements that should sit uniformly in front of or behind everything else (a translucent cartoon backdrop, a highlight band drawn boldly on top, etc.).A nested
list[PymolConfig]packs all the included layers into one shared PyMOL scene and a singlecmd.ray()call. PyMOL’s z-buffer interleaves their geometries per pixel (like what PyMol’s nativesupercommand does for two superposed structures) and each layer’stransparencybecomes a depth-aware per-selection setting (cartoon_transparency,transparencyfor surfaces,stick_transparency, etc.).
Example:
protein_config = portein.ProteinConfig(
pdb_file="../_data/7lc2.pdb",
rotate=True,
width=600,
chain_colormap="Set3",
output_prefix=str(output_dir / "compositing_demo"),
)
# Two bare layers — each is its own ray-trace, then PIL-composited.
# The cartoon PNG renders on top of the translucent surface PNG.
protein_config.output_prefix = str(output_dir / "decals")
decals_image = portein.Pymol(
protein=protein_config,
layers=[
portein.PymolConfig(representation="surface", pymol_settings=pymol_settings, transparency=0.5),
portein.PymolConfig(representation="cartoon", pymol_settings=pymol_settings),
],
).run()
portein.crop_to_content(Image.open(decals_image))
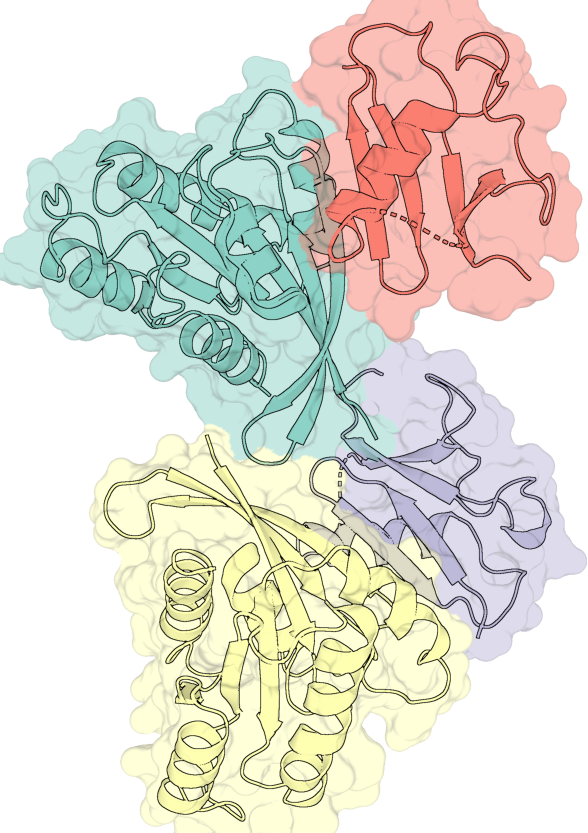
# One nested-list group — both layers share the same ray-trace.
# The translucent surface and the cartoon share the z-buffer: the cartoon
# appears behind wherever the surface is in front, and
# unmuted where the cartoon protrudes.
protein_config.output_prefix = str(output_dir / "grouped")
grouped_image = portein.Pymol(
protein=protein_config,
layers=[
[
portein.PymolConfig(representation="surface", pymol_settings=pymol_settings, transparency=0.5),
portein.PymolConfig(representation="cartoon", pymol_settings=pymol_settings),
],
],
).run()
portein.crop_to_content(Image.open(grouped_image))
When to use which:
Want |
Layer entry |
|---|---|
Uniform 2D translucency over each whole layer |
bare |
Depth-correct interleaving of overlapping geometries |
|
Two superposed structures rendered like PyMOL’s |
one nested group |
Surface + sticks where the ligand should occlude correctly |
one nested group |
Highlight band drawn boldly on top regardless of depth |
bare |
You can mix the two in one render — for example, a translucent cartoon as a backdrop (bare) plus a nested group of side-chain + ligand sticks that need to weave through each other:
layers=[
portein.PymolConfig(representation="cartoon", selection="all", transparency=0.5),
[
portein.PymolConfig(representation="sticks", selection="resi 14+15+16"),
portein.PymolConfig(representation="sticks", selection="resn GNP", color="green"),
],
]
From the command line#
Save a YAML with the protein config:
pdb_file: 7lc2
rotate: true
width: 1000
chain_colormap: Set3
output_prefix: examples/7lc2_simple
Then run:
portein pymol protein.yaml
portein pymol protein.yaml pymol_layers.yaml --buffer 10